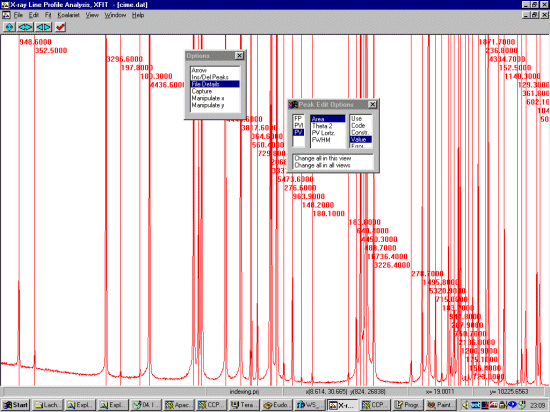
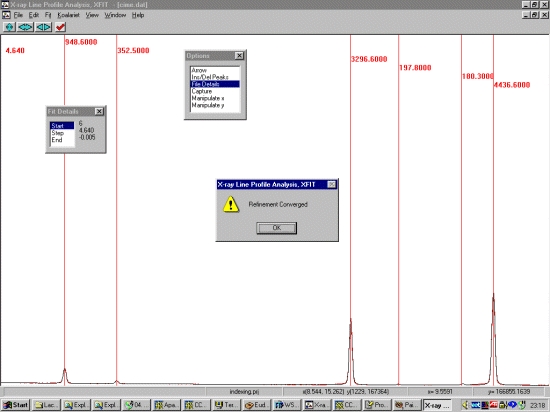
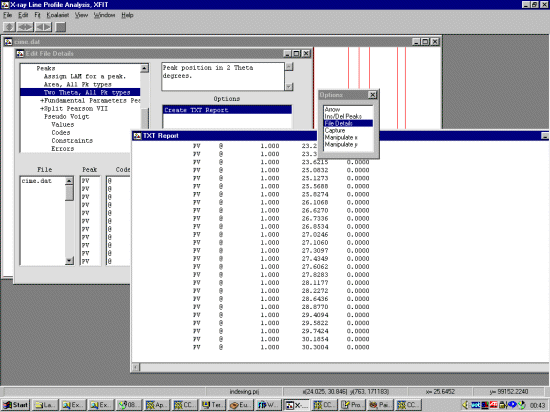
Open up the file and browse the pattern noting the range which gives fairly resolved peaks for profiling. At least 20 peaks are required. Using the File, Details option, set the background to refine with a bit more flexibility (3rd to 5th order).
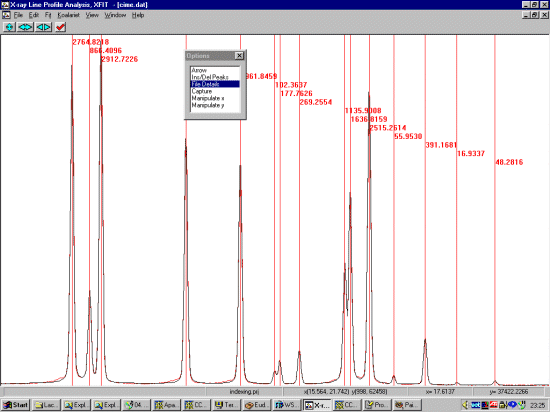
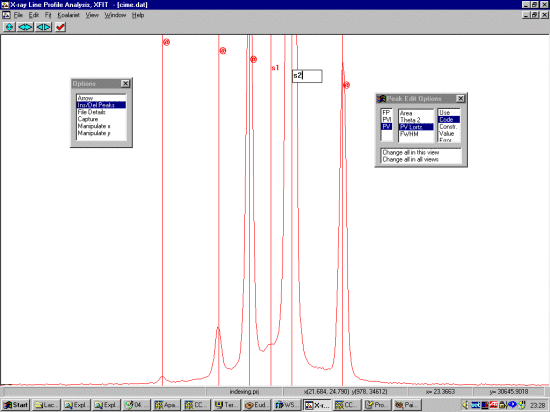
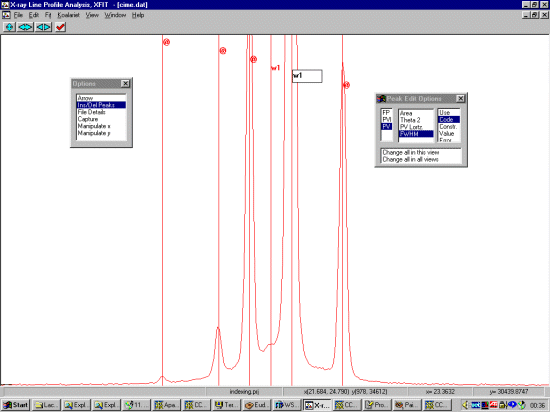
Selecting Ins/Del Peaks, then PV for Pseudo Voight, click left button on the peaks to profile. Right mouse button to remove a peak. If there are subtle shoulders or humps you are not sure about, ignore them for the moment. Come back to the shoulders if a single Pseudo-Voight peak cannot fit them. Also, if the width of a peak is enlarged compared to its neighbours, this could imply there are really two peaks.
In the following screen dump, we have selected our range.