
Site map









Site map |
||||
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
By means of the control file for the plaster sample
(plaster.sav) the necessary elements will be explained here.
Please open BGMNwin and select
File→Open Control File,
browse to the PLASTER\plaster.sav file and open it:
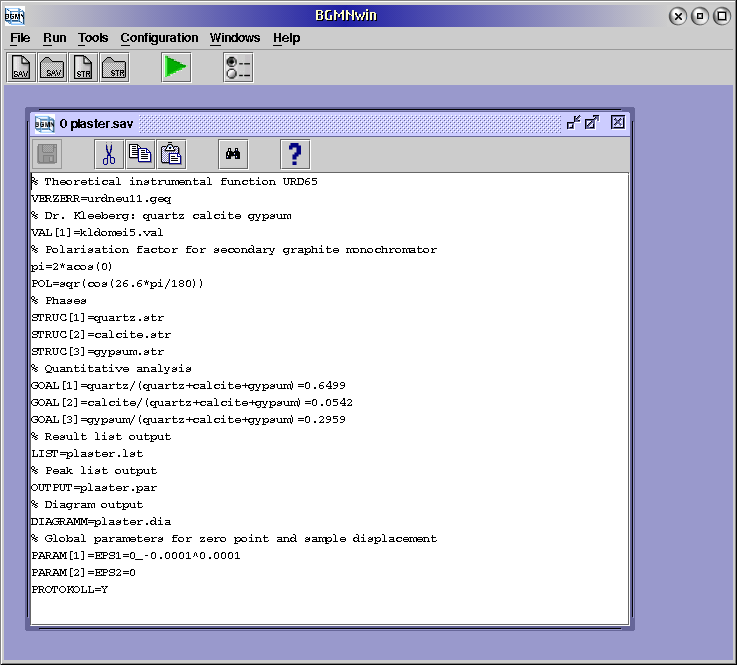
For an extended explanation of a variable put the cursor of your BGMNwin window to that variable and press F1 or see a list of variables.
First, the instrumental function has to be provided:
VERZERR=urdneu11.geqThen the pattern data file is needed:
VAL[1]=kldomei5.val
This file is in the APX-63 format, a simple text format from the SEIFERT-FPM diffractometers. Also, BGMN can process files with the extension 'rd' (Philips APD) and 'raw' (SIEMENS Diffrac-AT) and two variants of 'gsa' (GSAS) as well. If you want to use other formats, you have to convert it into one of the above.
If a secondary graphite monochromator was used, an appropriate polarisation factor is (26.6° is the 2Θ angle of the graphite crystal; without monochromator use 'POL=1', which is the default):
pi=2*acos(0) POL=sqr(cos(26.6*pi/180))The structure files are given in this way:
STRUC[1]=quartz.str STRUC[2]=calcite.str STRUC[3]=gypsum.strAnd the formulae for the quantitative analysis are:
GOAL[1]=quartz/(quartz+calcite+gypsum) GOAL[2]=calcite/(quartz+calcite+gypsum) GOAL[3]=gypsum/(quartz+calcite+gypsum)In this file (
plaster.lst) the results are written:
LIST=plaster.lstAn output file for the peak list (
plaster.par) is necessary if
you want a peak list:
OUTPUT=plaster.parFor the easy inspection of diagrams a file (
plaster.dia) with
pattern, calculated, difference and background data can be written:
DIAGRAMM=plaster.diaLast, the global parameters for the zero point (
EPS1) and sample
displacement (EPS2) are included:
PARAM[1]=EPS1=0_-0.0001^0.0001 PARAM[2]=EPS2=0Select
PROTOKOLL=Yto get more information about the refinement progress.
If you made any changes to plaster.sav, please save it.
The calculation is started by the command
Run→BGMN
Browse to the plaster.sav file and open it,
and after each iteration step a line with information about the iteration
number, the least squares sum and some other values is printed.
RESOUT[1]=quartz.res
PDBOUT[1]=quartz.pdb
STRUCOUT[1]=quartz.stoFor further output see Result Output